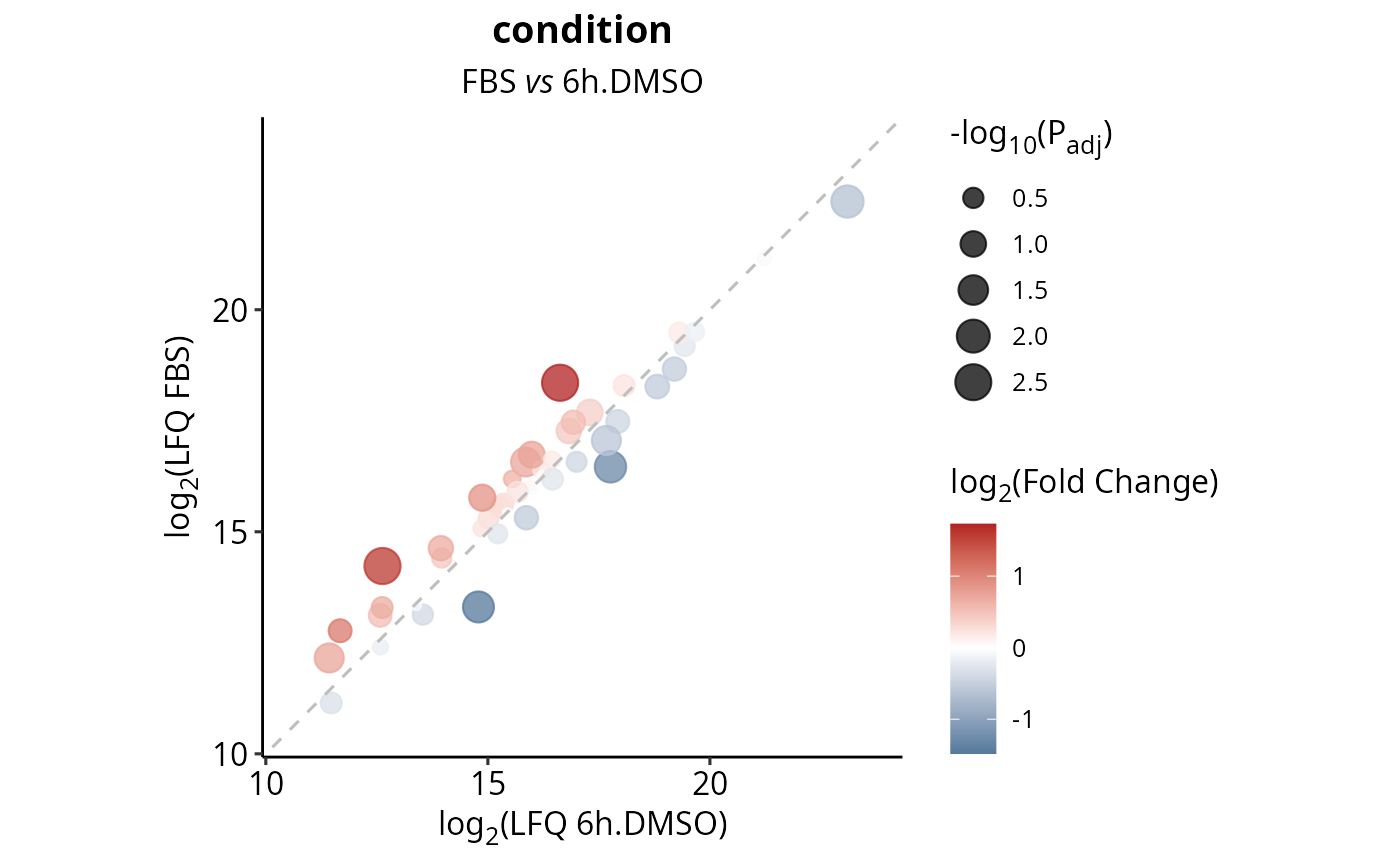
Plots a scatter of the average LFQ values of each condition in a contrast. Dots will be colored by FoldChange and the size will correspond to the -log10(Padj).
contrast.LFQ(
DEprot.analyses.object,
contrast = 1,
show.only.differential = FALSE,
show.only.significant = FALSE,
log2FC.positive.color = "firebrick",
log2FC.negative.color = "steelblue4",
log2FC.scale.min = NULL,
log2FC.scale.max = NULL,
identical.axes = TRUE,
dot.labels = NULL,
protein.names.pattern = "",
labels.in.boxes = FALSE,
label.font.size = 3,
label.max.overlaps = 100,
min.segment.length.labels = 0
)Arguments
- DEprot.analyses.object
An object of class
DEprot.analyses.- contrast
Integer indicating the position of the contrast to use. Default:
1.- show.only.differential
Logical value indicating whether only up or down regulated proteins should be plotted. Default:
FALSE.- show.only.significant
Logical value indicating whether only proteins with a significant p-value adjusted should be plotted. Notice that this is subordinated to `show.only.differential` (differential proteins pass the P-adj threshold by default). Default:
FALSE.- log2FC.positive.color
String indicating any R-supported color for the positive foldchange values. Default:
"firebrick".- log2FC.negative.color
String indicating any R-supported color for the negative foldchange values. Default:
"steelblue4".- log2FC.scale.min
Numeric value indicating the minimum of the foldchange scale (lower values will be colored with the minimum of the scale). Default:
NULL.- log2FC.scale.max
Numeric value indicating the maximum of the foldchange scale (lower values will be colored with the maximum of the scale). Default:
NULL.- identical.axes
Logical value indicating whether the axes should be forces to be identical between x and y. Default:
TRUE.- dot.labels
String vector indicating labels to show on the plot that should correspond to
prot.idcolumn values. Default:NULL(no labels shown).- protein.names.pattern
Character indicating a regular expression to remove from the protein IDs. Default:
"", no alterations in the protein IDs.- labels.in.boxes
Logical value indicating whether the labels should be visualized as boxes. Default:
FALSE.- label.font.size
Numeric value indicating the size to use for the dot labels. Default:
3.- label.max.overlaps
Numeric value indicating the maximum number of overlaps allowed between labels. Default:
100.- min.segment.length.labels
Numeric value indicating the minimal length of the segments that connect the labels to the points. Default:
0(segment always shown).
Value
A scatter plot of class ggplot2.
Examples
contrast.LFQ(DEprot.analyses.object = DEprot::test.toolbox$diff.exp.limma,
contrast = 1)