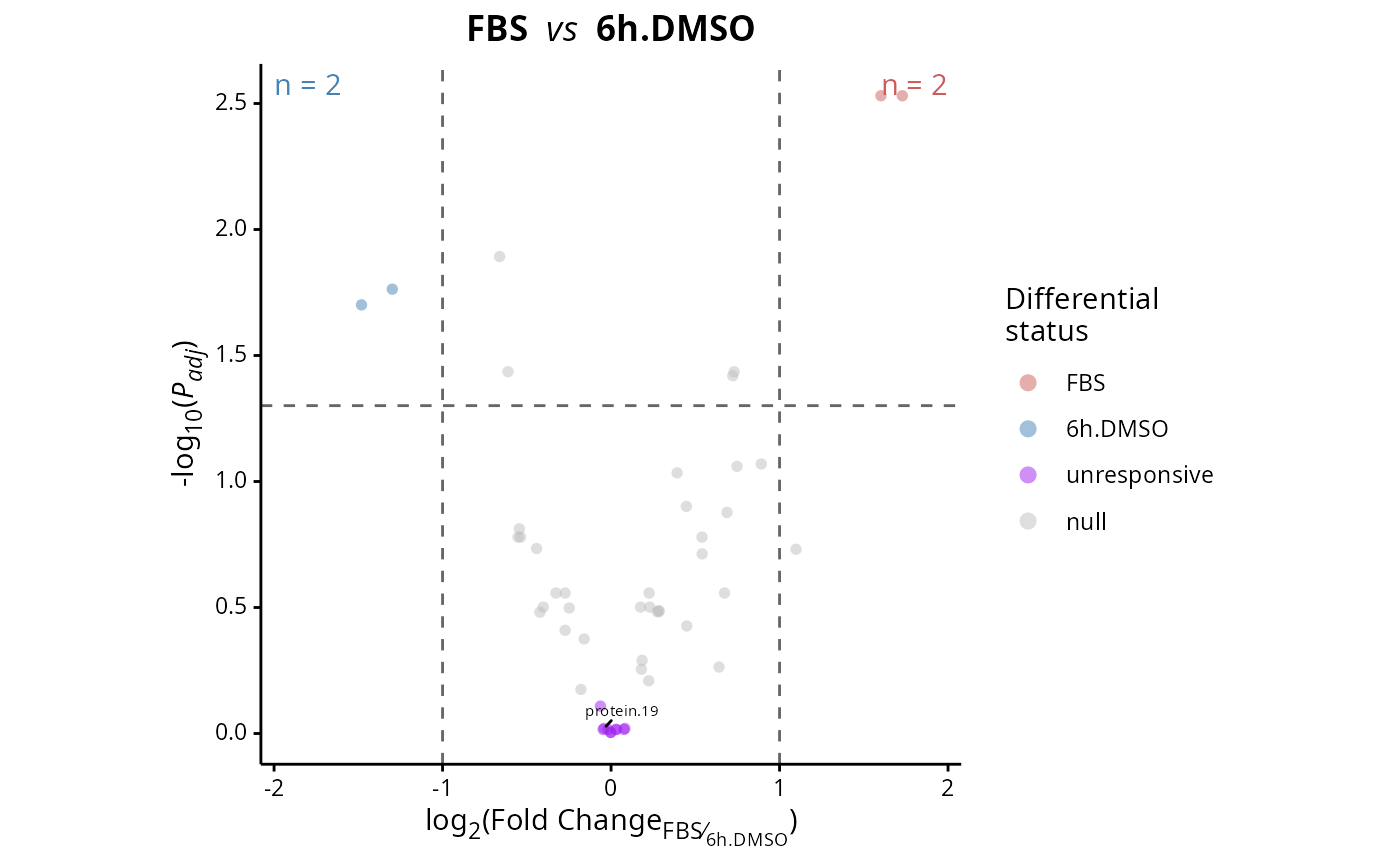
Plots a volcano plot log2(FoldChange) x -log10(p.adjusted) of differential expression results
# S3 method for class 'volcano'
plot(
DEprot.analyses.object,
contrast = 1,
up.color = "indianred",
down.color = "steelblue",
unresponsive.color = "purple",
null.color = "gray",
point.size = 2,
point.alpha = 0.5,
title = NULL,
use.uncorrected.pvalue = FALSE,
symmetric.x = TRUE,
dot.labels = NULL,
protein.names.pattern = "",
labels.in.boxes = FALSE,
label.font.size = 2,
label.max.overlaps = 100,
min.segment.length.labels = 0
)Arguments
- DEprot.analyses.object
An object of class
DEprot.analyses.- contrast
Number indicating the position of the contrast to use for the plotting.
- up.color
String indicating the color to use for up-regulated proteins in the plots. Default:
"indianred".- down.color
String indicating the color to use for up-regulated proteins in the plots. Default:
"steelblue".- unresponsive.color
String indicating the color to use for unresponsive proteins in the plots. Default:
"purple".- null.color
String indicating the color to use for null proteins in the plots. Default:
"gray".- point.size
Numeric value indicating the size of the dots. Default:
2.- point.alpha
Numeric value between 0 and 1 to indicate the transparency (alpha) of the dots. Default:
0.5.- title
String indicating the title to use. Default:
NULL(automatic title).- use.uncorrected.pvalue
Logical value indicating whether it should be used the normal p-value instead of the adjusted one (differential proteins numbers are recomputed). Default:
FALSE, padj is used.- symmetric.x
Logical values indicating whether the x-axis scale should be symmetric or not. Default:
TRUE.- dot.labels
String vector indicating labels to show on the plot that should correspond to
prot.idcolumn values. Default:NULL(no labels shown).- protein.names.pattern
Character indicating a regular expression to remove from the protein IDs. Default:
"", no alterations in the protein IDs.- labels.in.boxes
Logical value indicating whether the labels should be visualized as boxes. Default:
FALSE.- label.font.size
Numeric value indicating the size to use for the dot labels. Default:
2.- label.max.overlaps
Numeric value indicating the maximum number of overlaps allowed between labels. Default:
100.- min.segment.length.labels
Numeric value indicating the minimal length of the segments that connect the labels to the points. Default:
0(segment always shown).
Value
A ggplot object.
Examples
plot.volcano(DEprot.analyses.object = DEprot::test.toolbox$diff.exp.limma,
contrast = 1,
dot.labels = "protein.19",
labels.in.boxes = FALSE) +
ggplot2::xlab("log<sub>2</sub>(Fold Change FBS/6h.DMSO)")